Viljakainen L, Fürst MA, Grasse AV, Jurvansuu J, Oh J, Tolonen L, Eder T, Rattei T, Cremer S
Front. Microbiol. 14:1119002
doi: 10.3389/fmicb.2023.1119002
Abstract
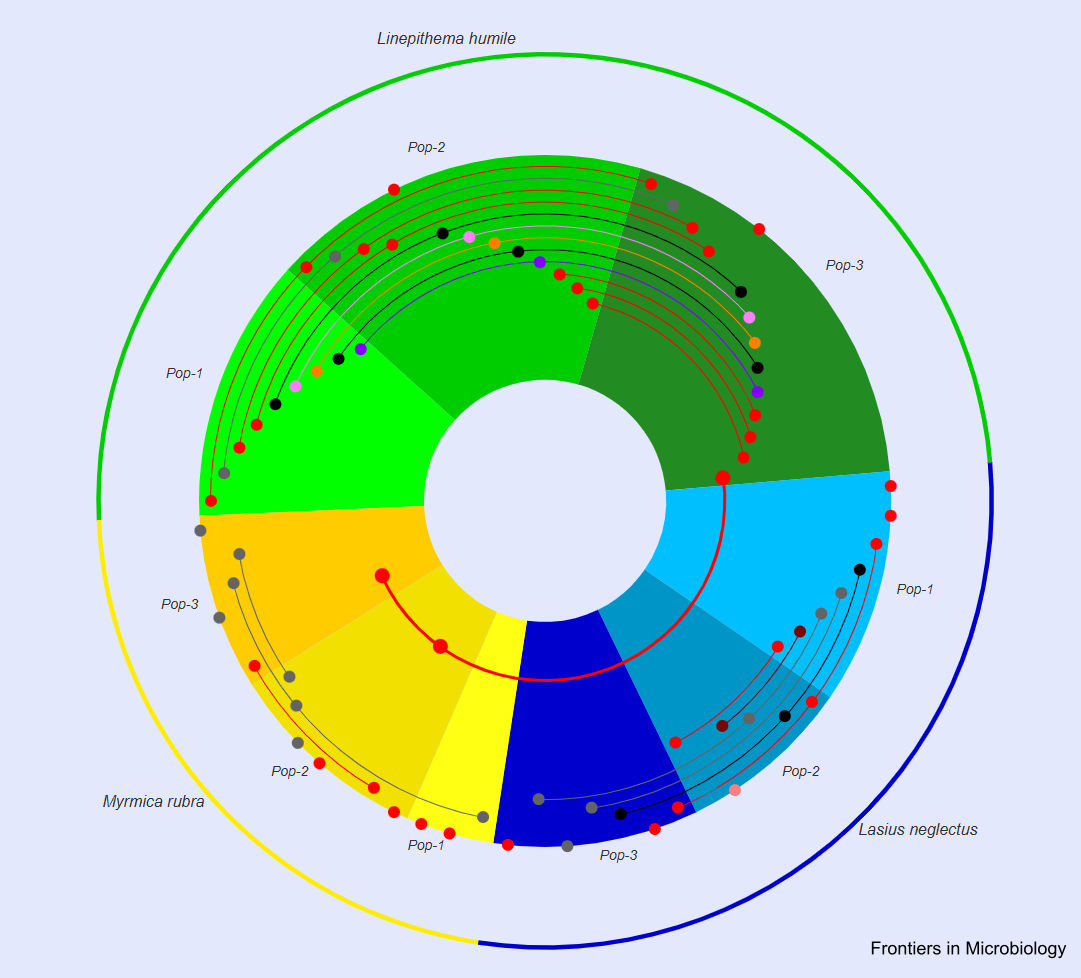
Hosts can carry many viruses in their bodies, but not all of them cause disease. We studied ants as a social host to determine both their overall viral repertoire and the subset of actively infecting viruses across natural populations of three subfamilies: the Argentine ant (Linepithema humile, Dolichoderinae), the invasive garden ant (Lasius neglectus, Formicinae) and the red ant (Myrmica rubra, Myrmicinae). We used a dual sequencing strategy to reconstruct complete virus genomes by RNA-seq and to simultaneously determine the small interfering RNAs (siRNAs) by small RNA sequencing (sRNA-seq), which constitute the host antiviral RNAi immune response. This approach led to the discovery of 41 novel viruses in ants and revealed a host ant-specific RNAi response (21 vs. 22 nt siRNAs) in the different ant species. The efficiency of the RNAi response (sRNA/RNA read count ratio) depended on the virus and the respective ant species, but not its population. Overall, we found the highest virus abundance and diversity per population in Li. humile, followed by La. neglectus and M. rubra. Argentine ants also shared a high proportion of viruses between populations, whilst overlap was nearly absent in M. rubra. Only one of the 59 viruses was found to infect two of the ant species as hosts, revealing high host-specificity in active infections. In contrast, six viruses actively infected one ant species, but were found as contaminants only in the others. Disentangling spillover of disease-causing infection from non-infecting contamination across species is providing relevant information for disease ecology and ecosystem management.